 This process is also used by molecular biologists (biologists whose studies involve DNA, RNA, and proteins) to create something called a cDNA library. They reverse-transcribe their RNA and incorporate it into the DNA of the host cell. What you just did in Table 3 is referred to as “reverse transcription.” This is actually what some RNA viruses do when they infect cells. 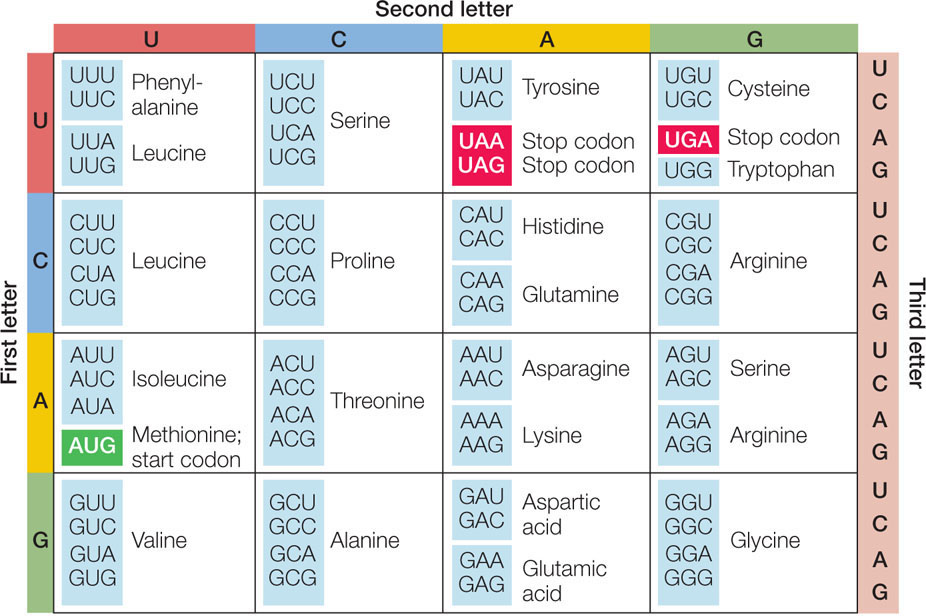
Table 3: Determining a DNA sequence that is complementary to the given piece of RNA and would code for the production of the DDG portion of a CESA protein. Remembering that adenine is complementary to thymine and uracil, and cytosine is complementary to guanine, fill in the sequences of complementary deoxyribonucletides that would have guided the production of this mRNA in the process of transcription. It also shows one of the possible RNA sequences that would be translated as that amino acid sequence. Table 3 shows a sequence of amino acids that makes up one of these conserved regions of a CESA protein. Transcription rewrites the information found in the sequence of deoxyribonucleotides into ribonucleotides. Remember, these codons are found on mRNA, and mRNA is created through the process of transcription. List all possible codons for these amino acids: Table 2: Possible codons which could product the indicated amino acids. Use the codon chart in Figure 2 to determine all the possible codons that would result in the incorporation of the amino acids indicated previously in Table 1. Since there are only 20 different types of amino acids that get incorporated into proteins, some amino acids can be coded for by multiple codons. Since codons consist of a sequence of three nucleotides and there are four different nucleotides in RNA and DNA, there are 64 different possible codons. This is what gives each amino acid its distinct properties. Notice that the side chain differs on each amino acid. The full name, three-letter and 1-letter abbreviations are given for each. 
Use the information in Figure 1 and write the full name for each of these amino acids.įigure 1: These are the amino acids incorporated into proteins. Table 1: One of the conserved amino acid sequences coded by all cellulose synthase genes is DDG. Use the chart in Figure 1 and write the names of the amino acids in the sequence DDG. For example, all the CesA genes have a region that codes for the following sequence of amino acids (each letter represents a certain amino acid sequence): DDG. They all have DNA sequences with “ conserved regions”, meaning that the code produces proteins that have sections where the amino acid sequences are identical. The genes in all cellulose producing plants studied so far have some things in common. The genes that code for the proteins that make cellulose are called CesA genes, and they were first identified from a bacterium that makes cellulose, Acetobacter xylinus (Saxena, Lin, and Brown, 1990). Even now, as they expand their studies to include additional species of plants and algae, this is where they start.Ĭellulose is an important part of the cell walls of plants, most algae, and even some prokaryotes. So, when the Roberts lab began studying the evolution of cellulose synthesis, one place they looked was at the available DNA sequence information. One way to study those roots is to look to DNA sequences. Cellulose Synthase Genes: Where do we start?Īll branches on the tree of life share common roots. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
